Régression logistique en 2D#
Links: notebook, html, PDF, python, slides, GitHub
Prédire la couleur d’un vin à partir de ses composants.
%matplotlib inline
from papierstat.datasets import load_wines_dataset
data = load_wines_dataset()
X = data.drop(['quality', 'color'], axis=1)
y = data['color']
from sklearn.model_selection import train_test_split
X_train, X_test, y_train, y_test = train_test_split(X, y)
from statsmodels.discrete.discrete_model import Logit
model = Logit(y_train == "white", X_train)
res = model.fit()
Optimization terminated successfully.
Current function value: 0.044575
Iterations 11
res.summary2()
| Model: | Logit | No. Iterations: | 11.0000 |
| Dependent Variable: | color | Pseudo R-squared: | 0.921 |
| Date: | 2018-02-08 00:13 | AIC: | 456.3398 |
| No. Observations: | 4872 | BIC: | 527.7436 |
| Df Model: | 10 | Log-Likelihood: | -217.17 |
| Df Residuals: | 4861 | LL-Null: | -2748.5 |
| Converged: | 1.0000 | Scale: | 1.0000 |
| Coef. | Std.Err. | z | P>|z| | [0.025 | 0.975] | |
|---|---|---|---|---|---|---|
| fixed_acidity | -1.4449 | 0.1519 | -9.5141 | 0.0000 | -1.7425 | -1.1472 |
| volatile_acidity | -12.0133 | 0.9940 | -12.0857 | 0.0000 | -13.9615 | -10.0651 |
| citric_acid | 0.1218 | 1.1387 | 0.1070 | 0.9148 | -2.1101 | 2.3537 |
| residual_sugar | 0.0780 | 0.0578 | 1.3484 | 0.1775 | -0.0354 | 0.1913 |
| chlorides | -33.6942 | 4.1533 | -8.1125 | 0.0000 | -41.8346 | -25.5538 |
| free_sulfur_dioxide | -0.0474 | 0.0149 | -3.1804 | 0.0015 | -0.0767 | -0.0182 |
| total_sulfur_dioxide | 0.0691 | 0.0055 | 12.6272 | 0.0000 | 0.0584 | 0.0798 |
| density | 45.6528 | 4.4627 | 10.2299 | 0.0000 | 36.9061 | 54.3994 |
| pH | -10.0587 | 1.0255 | -9.8086 | 0.0000 | -12.0687 | -8.0488 |
| sulphates | -9.1971 | 1.0697 | -8.5980 | 0.0000 | -11.2936 | -7.1006 |
| alcohol | 0.5328 | 0.1284 | 4.1494 | 0.0000 | 0.2811 | 0.7844 |
On ne garde que les deux premières.
X_train2 = X_train.iloc[:, :2]
import pandas
df = pandas.DataFrame(X_train2.copy())
df['y'] = y_train
import matplotlib.pyplot as plt
fig, ax = plt.subplots(1, 1, figsize=(4, 4))
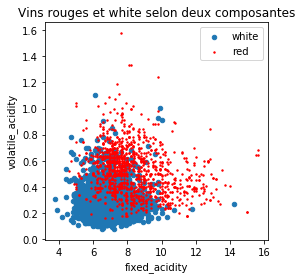
df[df.y == "white"].plot(x="fixed_acidity", y="volatile_acidity", ax=ax, kind='scatter', label="white")
df[df.y == "red"].plot(x="fixed_acidity", y="volatile_acidity", ax=ax,
kind='scatter', label="red", color="red", s=2)
ax.set_title("Vins rouges et white selon deux composantes");
from sklearn.linear_model import LogisticRegression
model = LogisticRegression()
model.fit(X_train2, y_train == "white")
LogisticRegression(C=1.0, class_weight=None, dual=False, fit_intercept=True,
intercept_scaling=1, max_iter=100, multi_class='ovr', n_jobs=1,
penalty='l2', random_state=None, solver='liblinear', tol=0.0001,
verbose=0, warm_start=False)
model.coef_, model.intercept_
(array([[ -0.92015604, -10.76494765]]), array([11.96976599]))
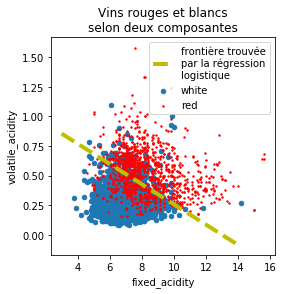
On trace cette droite sur le graphique.
x0 = 3
y0 = -(model.coef_[0,0] * x0 + model.intercept_) / model.coef_[0,1]
x1 = 14
y1 = -(model.coef_[0,0] * x1 + model.intercept_) / model.coef_[0,1]
x0, y0, x1, y1
(3, array([0.85548933]), 14, array([-0.08475829]))
import matplotlib.pyplot as plt
fig, ax = plt.subplots(1, 1, figsize=(4, 4))
df[df.y == "white"].plot(x="fixed_acidity", y="volatile_acidity", ax=ax, kind='scatter', label="white")
df[df.y == "red"].plot(x="fixed_acidity", y="volatile_acidity", ax=ax,
kind='scatter', label="red", color="red", s=2)
ax.plot([x0, x1], [y0, y1], 'y--', lw=4, label='frontière trouvée\npar la régression\nlogistique')
ax.legend()
ax.set_title("Vins rouges et blancs\nselon deux composantes");