Note
Click here to download the full example code
Measuring CPU performance with a parallelized vector sum#
The example compares the time spend in computing the sum of all coefficients of a matrix when the function walks through the coefficients by rows or by columns when the computation is parallelized.
Vector Sum#
from tqdm import tqdm
import numpy
import matplotlib.pyplot as plt
from pandas import DataFrame
from onnx_extended.ext_test_case import measure_time, unit_test_going
from onnx_extended.validation._validation import (
vector_sum_array as vector_sum,
vector_sum_array_parallel as vector_sum_parallel,
)
obs = []
dims = [500, 700, 800, 900, 1000, 1100, 1200, 1300, 1400, 1500, 1600, 1700, 1800, 2000]
if unit_test_going():
dims = dims[:3]
for dim in tqdm(dims):
values = numpy.ones((dim, dim), dtype=numpy.float32).ravel()
diff = abs(vector_sum(dim, values, True) - dim**2)
res = measure_time(lambda: vector_sum(dim, values, True), max_time=0.5)
obs.append(
dict(
dim=dim,
size=values.size,
time=res["average"],
direction="rows",
time_per_element=res["average"] / dim**2,
diff=diff,
)
)
res = measure_time(lambda: vector_sum_parallel(dim, values, True), max_time=0.5)
obs.append(
dict(
dim=dim,
size=values.size,
time=res["average"],
direction="rows//",
time_per_element=res["average"] / dim**2,
diff=diff,
)
)
diff = abs(vector_sum(dim, values, False) - dim**2)
res = measure_time(lambda: vector_sum_parallel(dim, values, False), max_time=0.5)
obs.append(
dict(
dim=dim,
size=values.size,
time=res["average"],
direction="cols//",
time_per_element=res["average"] / dim**2,
diff=diff,
)
)
df = DataFrame(obs)
piv = df.pivot(index="dim", columns="direction", values="time_per_element")
print(piv)
0%| | 0/14 [00:00<?, ?it/s]
7%|7 | 1/14 [00:01<00:22, 1.72s/it]
14%|#4 | 2/14 [00:03<00:20, 1.72s/it]
21%|##1 | 3/14 [00:05<00:19, 1.74s/it]
29%|##8 | 4/14 [00:07<00:17, 1.78s/it]
36%|###5 | 5/14 [00:08<00:16, 1.83s/it]
43%|####2 | 6/14 [00:10<00:14, 1.85s/it]
50%|##### | 7/14 [00:12<00:12, 1.82s/it]
57%|#####7 | 8/14 [00:14<00:10, 1.81s/it]
64%|######4 | 9/14 [00:16<00:09, 1.84s/it]
71%|#######1 | 10/14 [00:18<00:07, 1.87s/it]
79%|#######8 | 11/14 [00:20<00:05, 1.87s/it]
86%|########5 | 12/14 [00:22<00:03, 1.90s/it]
93%|#########2| 13/14 [00:24<00:01, 1.93s/it]
100%|##########| 14/14 [00:26<00:00, 1.94s/it]
100%|##########| 14/14 [00:26<00:00, 1.86s/it]
direction cols// rows rows//
dim
500 4.238436e-08 2.325730e-09 4.092764e-08
700 2.157749e-08 2.232951e-09 2.038646e-08
800 1.823866e-08 2.212191e-09 1.633296e-08
900 1.416694e-08 2.189656e-09 1.278406e-08
1000 1.104241e-08 2.171995e-09 1.017222e-08
1100 9.759492e-09 2.179661e-09 8.933916e-09
1200 7.967947e-09 2.194684e-09 7.480013e-09
1300 8.483947e-09 2.175047e-09 4.195949e-09
1400 6.115952e-09 2.185732e-09 4.635022e-09
1500 6.646210e-09 2.167680e-09 4.261122e-09
1600 5.386631e-09 2.127125e-09 3.333226e-09
1700 4.713320e-09 2.144145e-09 2.616001e-09
1800 5.660132e-09 2.172713e-09 3.167737e-09
2000 4.173346e-09 2.117624e-09 2.657948e-09
Plots#
piv_diff = df.pivot(index="dim", columns="direction", values="diff")
piv_time = df.pivot(index="dim", columns="direction", values="time")
fig, ax = plt.subplots(1, 3, figsize=(12, 6))
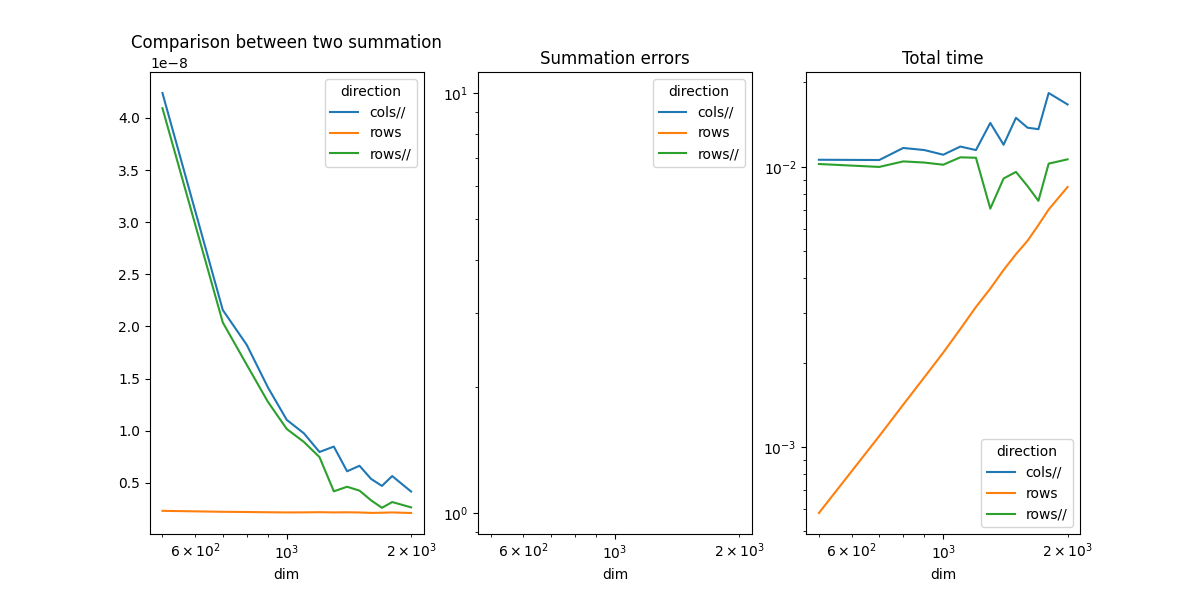
piv.plot(ax=ax[0], logx=True, title="Comparison between two summation")
piv_diff.plot(ax=ax[1], logx=True, logy=True, title="Summation errors")
piv_time.plot(ax=ax[2], logx=True, logy=True, title="Total time")
fig.savefig("plot_bench_cpu_vector_sum_parallel.png")
/usr/local/lib/python3.9/site-packages/pandas/plotting/_matplotlib/core.py:744: UserWarning: Data has no positive values, and therefore cannot be log-scaled.
labels = axis.get_majorticklabels() + axis.get_minorticklabels()
The summation by rows is much faster as expected. That explains why it is usually more efficient to transpose the first matrix before a matrix multiplication. Parallelization is faster.
Total running time of the script: ( 0 minutes 31.100 seconds)