Prédiction de la note des vins#
Links: notebook, html, PDF, python, slides, GitHub
Le notebook compare plusieurs de modèles de régression.
%matplotlib inline
import warnings
warnings.simplefilter('ignore')
from papierstat.datasets import load_wines_dataset
df = load_wines_dataset()
X = df.drop(['quality', 'color'], axis=1)
y = yn = df['quality']
from sklearn.model_selection import train_test_split
X_train, X_test, y_train, y_test = train_test_split(X, y)
On normalise les données.
from sklearn.preprocessing import normalize
X_train_norm = normalize(X_train)
X_test_norm = normalize(X_test)
X_norm = normalize(X)
Cette façon de faire est complètement erronnée car il est peu probable que la même normalisation soit apppliquée sur les trois bases. La normalisation doit être estimée sur la base d’apprentissage et appliquée sur la base de test. Reprenons.
from sklearn.preprocessing import Normalizer
norm = Normalizer()
X_train_norm = norm.fit_transform(X_train)
X_test_norm = norm.transform(X_test)
X_norm = norm.transform(X)
On récupère beaucoup de modèles implémentés dans scikit-learn pour la régression.
from sklearn.linear_model import *
from sklearn.ensemble import *
from sklearn.neighbors import *
from sklearn.neural_network import *
models = [
model for name, model in globals().items() if (
hasattr(model, 'fit') and not hasattr(model, 'predict_proba') and
hasattr(model, '__name__') and not model.__name__.endswith('CV') and
'Logistic' not in model.__name__ and 'Regr' in model.__name__)]
import pprint
pprint.pprint(models)
[<class 'sklearn.linear_model._bayes.ARDRegression'>,
<class 'sklearn.linear_model._huber.HuberRegressor'>,
<class 'sklearn.linear_model._base.LinearRegression'>,
<class 'sklearn.linear_model._passive_aggressive.PassiveAggressiveRegressor'>,
<class 'sklearn.linear_model._quantile.QuantileRegressor'>,
<class 'sklearn.linear_model._stochastic_gradient.SGDRegressor'>,
<class 'sklearn.linear_model._theil_sen.TheilSenRegressor'>,
<class 'sklearn.linear_model._ransac.RANSACRegressor'>,
<class 'sklearn.linear_model._glm.glm.PoissonRegressor'>,
<class 'sklearn.linear_model._glm.glm.GammaRegressor'>,
<class 'sklearn.linear_model._glm.glm.TweedieRegressor'>,
<class 'sklearn.ensemble._forest.RandomForestRegressor'>,
<class 'sklearn.ensemble._forest.ExtraTreesRegressor'>,
<class 'sklearn.ensemble._bagging.BaggingRegressor'>,
<class 'sklearn.ensemble._gb.GradientBoostingRegressor'>,
<class 'sklearn.ensemble._weight_boosting.AdaBoostRegressor'>,
<class 'sklearn.ensemble._voting.VotingRegressor'>,
<class 'sklearn.ensemble._stacking.StackingRegressor'>,
<class 'sklearn.ensemble._hist_gradient_boosting.gradient_boosting.HistGradientBoostingRegressor'>,
<class 'sklearn.neighbors._regression.KNeighborsRegressor'>,
<class 'sklearn.neighbors._regression.RadiusNeighborsRegressor'>,
<class 'sklearn.neural_network._multilayer_perceptron.MLPRegressor'>]
from sklearn.metrics import r2_score
def score_model(xtr, xte, ytr, yte, model):
try:
model.fit(xtr, ytr)
except Exception as e:
raise Exception("Issue with model '{0}'".format(model.__name__)) from e
return r2_score(yte, model.predict(xte))
from time import perf_counter
r2s = []
names = []
durations = []
regressors = {}
for i, model in enumerate(models):
if model.__name__ in {'ARDRegression', 'VotingRegressor', 'StackingRegressor',
'QuantileRegressor'}:
continue
try:
reg = model()
except Exception as e:
print('Skip', model)
continue
begin = perf_counter()
r2 = score_model(X_train_norm, X_test_norm, y_train, y_test, reg)
duree = perf_counter () - begin
r2s.append(r2)
names.append(model.__name__)
durations.append(duree)
regressors[model.__name__] = reg
print(i, model.__name__, r2, duree)
1 HuberRegressor 0.18696322297535295 0.10263620000000007
2 LinearRegression 0.18670706392432423 0.0031078999999998302
3 PassiveAggressiveRegressor -2.4493503320572922 0.00904460000000018
5 SGDRegressor 0.014418866580130474 0.005636599999999881
6 TheilSenRegressor -0.22921086116095046 2.5755812999999996
7 RANSACRegressor -0.22456172730313217 0.10757049999999957
8 PoissonRegressor 0.0019476060926755245 0.008242899999999942
9 GammaRegressor 0.00035190831663889366 0.0046112000000002595
10 TweedieRegressor 0.00035356971012889815 0.00390669999999993
11 RandomForestRegressor 0.4902308886152582 3.781266499999999
12 ExtraTreesRegressor 0.5234939074444616 1.2828908000000006
13 BaggingRegressor 0.4275524601097942 0.3720353999999997
14 GradientBoostingRegressor 0.33578249602447263 2.3406590000000005
15 AdaBoostRegressor 0.19852494764300066 0.5364822
18 HistGradientBoostingRegressor 0.4116613218417975 1.8669652999999986
19 KNeighborsRegressor 0.17441896259812206 0.06659600000000054
20 RadiusNeighborsRegressor 0.00012107760711554949 0.7381427999999985
21 MLPRegressor 0.18147670458010268 4.3011114
import pandas
df = pandas.DataFrame(dict(model=names, r2=r2s, duree=durations))
df = df[['model', 'r2', 'duree']]
df.sort_values('r2')
| model | r2 | duree | |
|---|---|---|---|
| 2 | PassiveAggressiveRegressor | -2.449350 | 0.009045 |
| 4 | TheilSenRegressor | -0.229211 | 2.575581 |
| 5 | RANSACRegressor | -0.224562 | 0.107570 |
| 16 | RadiusNeighborsRegressor | 0.000121 | 0.738143 |
| 7 | GammaRegressor | 0.000352 | 0.004611 |
| 8 | TweedieRegressor | 0.000354 | 0.003907 |
| 6 | PoissonRegressor | 0.001948 | 0.008243 |
| 3 | SGDRegressor | 0.014419 | 0.005637 |
| 15 | KNeighborsRegressor | 0.174419 | 0.066596 |
| 17 | MLPRegressor | 0.181477 | 4.301111 |
| 1 | LinearRegression | 0.186707 | 0.003108 |
| 0 | HuberRegressor | 0.186963 | 0.102636 |
| 13 | AdaBoostRegressor | 0.198525 | 0.536482 |
| 12 | GradientBoostingRegressor | 0.335782 | 2.340659 |
| 14 | HistGradientBoostingRegressor | 0.411661 | 1.866965 |
| 11 | BaggingRegressor | 0.427552 | 0.372035 |
| 9 | RandomForestRegressor | 0.490231 | 3.781266 |
| 10 | ExtraTreesRegressor | 0.523494 | 1.282891 |
On filtre les valeurs inférieures à -1.
df = df[df.r2 >= -1]
import matplotlib.pyplot as plt
fig, ax = plt.subplots(figsize=(12, 4))
df.plot(x='duree', y='r2', kind="scatter", ax=ax)
for row in df.itertuples():
t, y, x = row[1:4]
if t[0] in {'B', 'L'}:
ax.text(x, y, t, rotation=30, ha='left', va='bottom')
else:
ax.text(x, y, t, rotation=-40)
ax.set_title("Comparaison des modèles selon le temps d'apprentissage / r2");
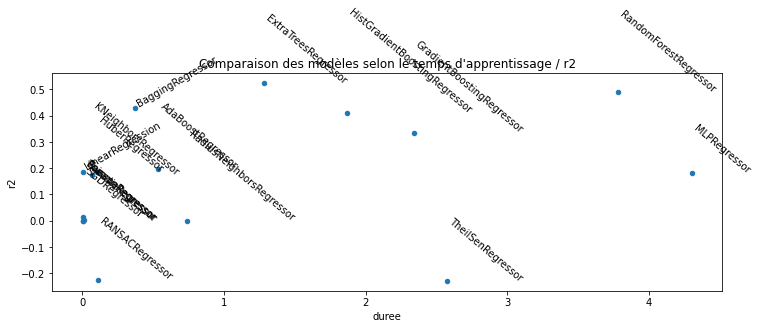
L’estimateur
RANSACRegressor
produit un très négatif. Regardons plus en détail.
pred = regressors['RANSACRegressor'].predict(X_test_norm)
import numpy.random
df = pandas.DataFrame(dict(pred=pred, expected=y_test))
df['expected'] += numpy.random.random(df.shape[0]) * 0.5
ax = df.plot(x="expected", y="pred", kind='scatter', figsize=(4, 4), linewidths=0.1)
ax.set_title('Graphe XY - valeurs attendues / prédites');
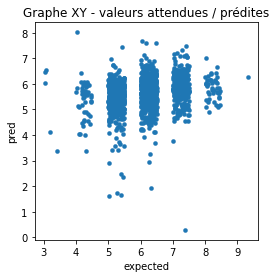
Essayons de voir avec la densité.
import seaborn
ax = seaborn.jointplot(df["expected"], df["pred"], kind="kde", size=4, space=0, ylim=(1, 9))
ax.ax_marg_y.set_title('Densité XY - valeur attendues / prédites');
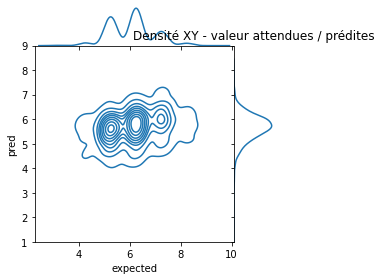
Pas facile à voir. Essayons de voir autrement en triant les prédictions et les valeurs attendues par ordre.
sv = df.sort_values(['expected', 'pred']).reset_index(drop=True)
import matplotlib.pyplot as plt
fig, ax = plt.subplots(figsize=(12, 3))
ax.plot(sv["pred"], label="RANSAC", lw=0.1)
ax.plot(list(sorted(df["expected"])), label="expected")
ax.plot(list(sorted(df["pred"])), label="RANSAC trié")
ax.set_ylim([0, 9])
ax.set_title('Valeur attendues et prédites triées par ordre croissant')
ax.legend();
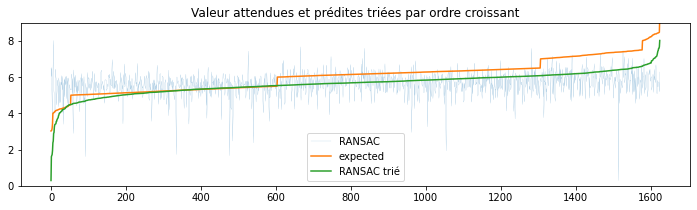
Le modèle est plutôt agité.
regressors['RANSACRegressor'].estimator_.intercept_
8.296147714878684
Pour s’assurer que les résultats sont fiables, il faut recommencer avec une validation croisée.
import numpy.random
rnd = numpy.random.permutation(range(X_norm.shape[0]))
xns = X_norm[rnd, :]
yns = yn[rnd]
xns.shape, yns.shape
((6497, 11), (6497,))
from sklearn.model_selection import cross_val_score
def score_model_cross(xn, yn, model):
res = cross_val_score(model, xn, yn, cv=5)
return res.mean(), min(res), max(res)
score_model_cross(xns, yns, LinearRegression())
(0.17975891799937466, 0.16696505316283117, 0.19857004070455042)
r2s = []
mis = []
mas = []
names = []
durations = []
regressors = {}
for i, model in enumerate(models):
if model.__name__ in {'ARDRegression', 'VotingRegressor', 'QuantileRegressor'}:
continue
try:
reg = model()
except Exception as e:
print('Skip', model)
continue
begin = perf_counter()
r2, mi, ma = score_model_cross(xns, yns, reg)
duree = perf_counter() - begin
r2s.append(r2)
mis.append(mi)
mas.append(ma)
names.append(model.__name__)
durations.append(duree)
regressors[model.__name__] = reg
print(i, model.__name__, r2, mi, ma, "t=%r" % duree)
1 HuberRegressor 0.15926614746076234 0.14654481316725898 0.16913851017974213 t=0.454639499999999
2 LinearRegression 0.17975891799937466 0.16696505316283117 0.19857004070455042 t=0.016710999999997256
3 PassiveAggressiveRegressor -1.2023166987836942 -3.9867877266892178 0.1473069141592367 t=0.060205800000002085
5 SGDRegressor 0.008672768127243647 0.0021660038805777493 0.019095545791309232 t=0.03673420000000149
6 TheilSenRegressor -0.36938713022242337 -0.6323773846900367 -0.1285090090856622 t=12.1253441
7 RANSACRegressor -1.216541764611399 -2.1838034440150125 -0.49602375704620316 t=0.4227926000000011
8 PoissonRegressor 0.0005335331789835696 -0.00324272848486995 0.002202890520605516 t=0.041724600000002
9 GammaRegressor -0.0011833153140562436 -0.005003239249327773 0.00038081596763739345 t=0.03832559999999319
10 TweedieRegressor -0.0011890320765876484 -0.005037356583908137 0.0003836189724416572 t=0.03498609999999758
11 RandomForestRegressor 0.49573724359207744 0.47719555839567485 0.5195191337192313 t=20.5839067
12 ExtraTreesRegressor 0.5205178519581393 0.5088671068579345 0.544928615205679 t=6.329529699999995
13 BaggingRegressor 0.44200753753008665 0.4148201125863905 0.4665893219045759 t=1.9267918999999978
14 GradientBoostingRegressor 0.33429915333938653 0.31585334825211187 0.34981047673804344 t=8.977681399999994
15 AdaBoostRegressor 0.2087316189255112 0.19118571603372658 0.22096772688033828 t=1.8667038999999903
Skip <class 'sklearn.ensemble._stacking.StackingRegressor'>
18 HistGradientBoostingRegressor 0.4206957666044492 0.3909890965983751 0.4324440282212091 t=4.953426099999987
19 KNeighborsRegressor 0.17992428041726455 0.14412273193552394 0.2237578623112627 t=0.3946573999999998
20 RadiusNeighborsRegressor -0.0014925783119640545 -0.005365612415162646 0.00012185111722451403 t=3.9200797999999963
21 MLPRegressor 0.18770733115748367 0.15333935822510514 0.21342290214312276 t=20.982291099999998
df = pandas.DataFrame(dict(model=names, r2=r2s, min=mis, max=mas, duree=durations))
df = df[['model', 'r2', 'min', 'max', 'duree']]
df.sort_values('r2')
| model | r2 | min | max | duree | |
|---|---|---|---|---|---|
| 5 | RANSACRegressor | -1.216542 | -2.183803 | -0.496024 | 0.422793 |
| 2 | PassiveAggressiveRegressor | -1.202317 | -3.986788 | 0.147307 | 0.060206 |
| 4 | TheilSenRegressor | -0.369387 | -0.632377 | -0.128509 | 12.125344 |
| 16 | RadiusNeighborsRegressor | -0.001493 | -0.005366 | 0.000122 | 3.920080 |
| 8 | TweedieRegressor | -0.001189 | -0.005037 | 0.000384 | 0.034986 |
| 7 | GammaRegressor | -0.001183 | -0.005003 | 0.000381 | 0.038326 |
| 6 | PoissonRegressor | 0.000534 | -0.003243 | 0.002203 | 0.041725 |
| 3 | SGDRegressor | 0.008673 | 0.002166 | 0.019096 | 0.036734 |
| 0 | HuberRegressor | 0.159266 | 0.146545 | 0.169139 | 0.454639 |
| 1 | LinearRegression | 0.179759 | 0.166965 | 0.198570 | 0.016711 |
| 15 | KNeighborsRegressor | 0.179924 | 0.144123 | 0.223758 | 0.394657 |
| 17 | MLPRegressor | 0.187707 | 0.153339 | 0.213423 | 20.982291 |
| 13 | AdaBoostRegressor | 0.208732 | 0.191186 | 0.220968 | 1.866704 |
| 12 | GradientBoostingRegressor | 0.334299 | 0.315853 | 0.349810 | 8.977681 |
| 14 | HistGradientBoostingRegressor | 0.420696 | 0.390989 | 0.432444 | 4.953426 |
| 11 | BaggingRegressor | 0.442008 | 0.414820 | 0.466589 | 1.926792 |
| 9 | RandomForestRegressor | 0.495737 | 0.477196 | 0.519519 | 20.583907 |
| 10 | ExtraTreesRegressor | 0.520518 | 0.508867 | 0.544929 | 6.329530 |
import matplotlib.pyplot as plt
fig, ax = plt.subplots(figsize=(12, 4))
df[df['min'] > -0.1].plot(x='duree', y='r2', kind="scatter", ax=ax)
for row in df.itertuples():
t, y, mi, ma, x = row[1:6]
if mi < -0.1:
continue
ax.plot([x, x], [mi, ma], color="black")
if t[0] in {'B', 'L'}:
ax.text(x, y, t, rotation=30, ha='left', va='bottom')
else:
ax.text(x, y, t, rotation=-40)
ax.set_ylim([-1, 1])
ax.set_title("Comparaison des modèles selon le temps d'apprentissage / r2\nValidation croisée");
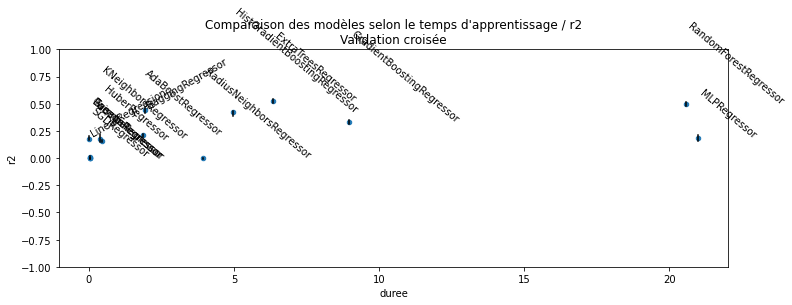

Le modèle RANSAC est conçu pour apprendre un modèle linéaire et réduire l’ensemble d’apprentissage aux points non aberrants. Dans notre cas, vu le peu d’exemples pour les notes élevées, il est très probable que celles-ci disparaissent des observations choisies pour estimer le modèle : le modèle choisit d’évincer les exemples pour lesquels l’erreur est la plus grande en considérant que cela est une indication du fait qu’ils sont aberrants. Malgré cela, sa faible capacité à prévoir vient du fait que la majorité des vins ont une note entre 4 et 6 et que finalement, il y a peu de différences : ils sont tous moyens et la différence s’explique par d’autres facteurs comme le juge ayant donné la note.